Why this case matters
About half of pediatric ulcerative colitis (UC) patients fail to achieve remission at week 4 on first-line corticosteroids or 5-ASA. Those non-responders accumulate cumulative steroid exposure, surgery risk, and growth impairment while the field waits for biologic-refractory rescue. The PROTECT inception cohort (GSE109142, 206 treatment-naïve pediatric UC patients with rectal biopsies at diagnosis and a clean week-4 remission readout) is one of the few datasets that lets us interrogate baseline mucosal biology before therapy obscures the signal.
We re-analysed PROTECT end-to-end with one question in mind: what druggable hypotheses survive rigorous causal-inference scrutiny, and which of those are actually translatable to a child? The answer reframes the repurposing landscape, surfaces a high-conviction novel target, and disqualifies an entire approved-drug class on developmental safety grounds.
A confounding-adjusted signature for an un-analysable cohort
Step 1: The confounding problem nobody can analyse away
PROTECT was designed for clinical care, not transcriptomics: 5-ASA was prescribed exclusively to low-severity patients (53/53), CS-IV concentrated in high-severity disease (47/72). The therapy × severity contingency table yields χ² = 139.88, p < 0.0001: near-perfect collinearity.
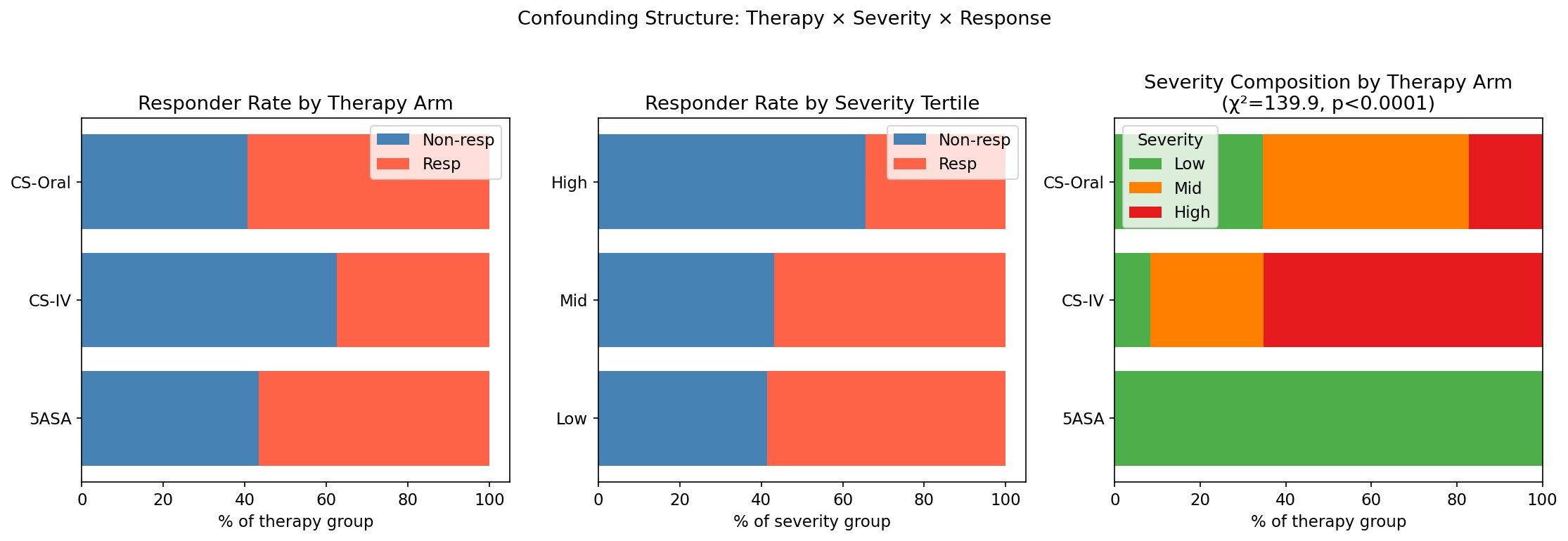
The consequence is brutal but informative. Across adjustment strategies, the gene-level signal collapses:
| Adjustment strategy | Genes at FDR < 0.05 | Median |logFC| |
|---|---|---|
| Naïve responder vs non-responder | hundreds | — |
| Severity-adjusted limma | 0 / 21,004 | 0.035 |
| Joint therapy + severity adjusted | 0 | 0.035 |
| Doubly-robust AIPW (IPTW + outcome) | 0 | 0.035 |
| E-value sensitivity (genome-wide median) | — | median E=1.19 |
Only 4 genes reach E ≥ 3 across the genome. The single naïvely-significant gene (BEAN1) attenuates to FDR 0.151 with E = 1.84. At the single-gene level, week-4 non-response is dominated by baseline disease severity, not a discoverable transcriptional driver. This is the kind of result that kills target-ID programmes, unless you escalate the analytical layer.
Step 2: The signal lives at the pathway, network, and TF level
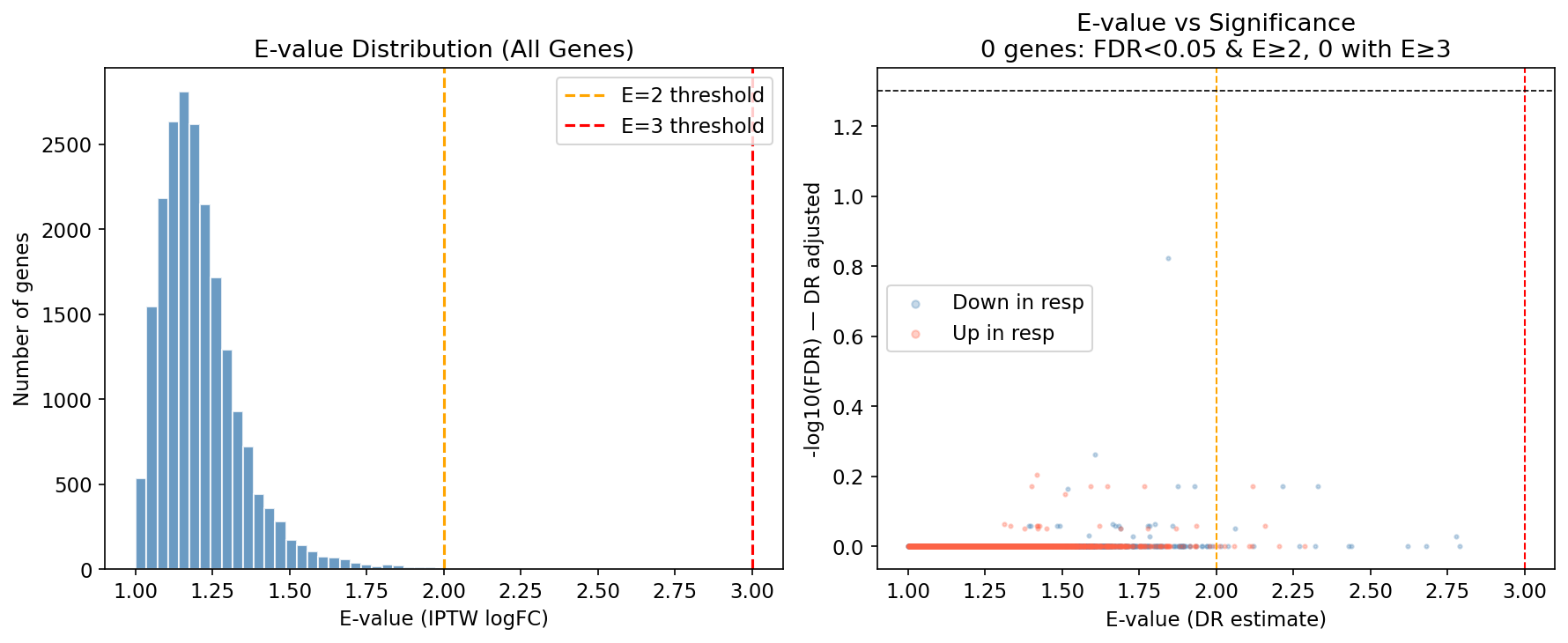
When the gene-level signal is confounding-bound, coordinated multi-gene programmes are not. Across GSEA, decoupleR TF activity (CollecTRI), WGCNA modules, and immune deconvolution we recovered 688 concordant pathways (FDR ≤ 0.25 in both naïve DE and IPTW rankings, zero direction-discordant) and a coherent two-programme architecture.
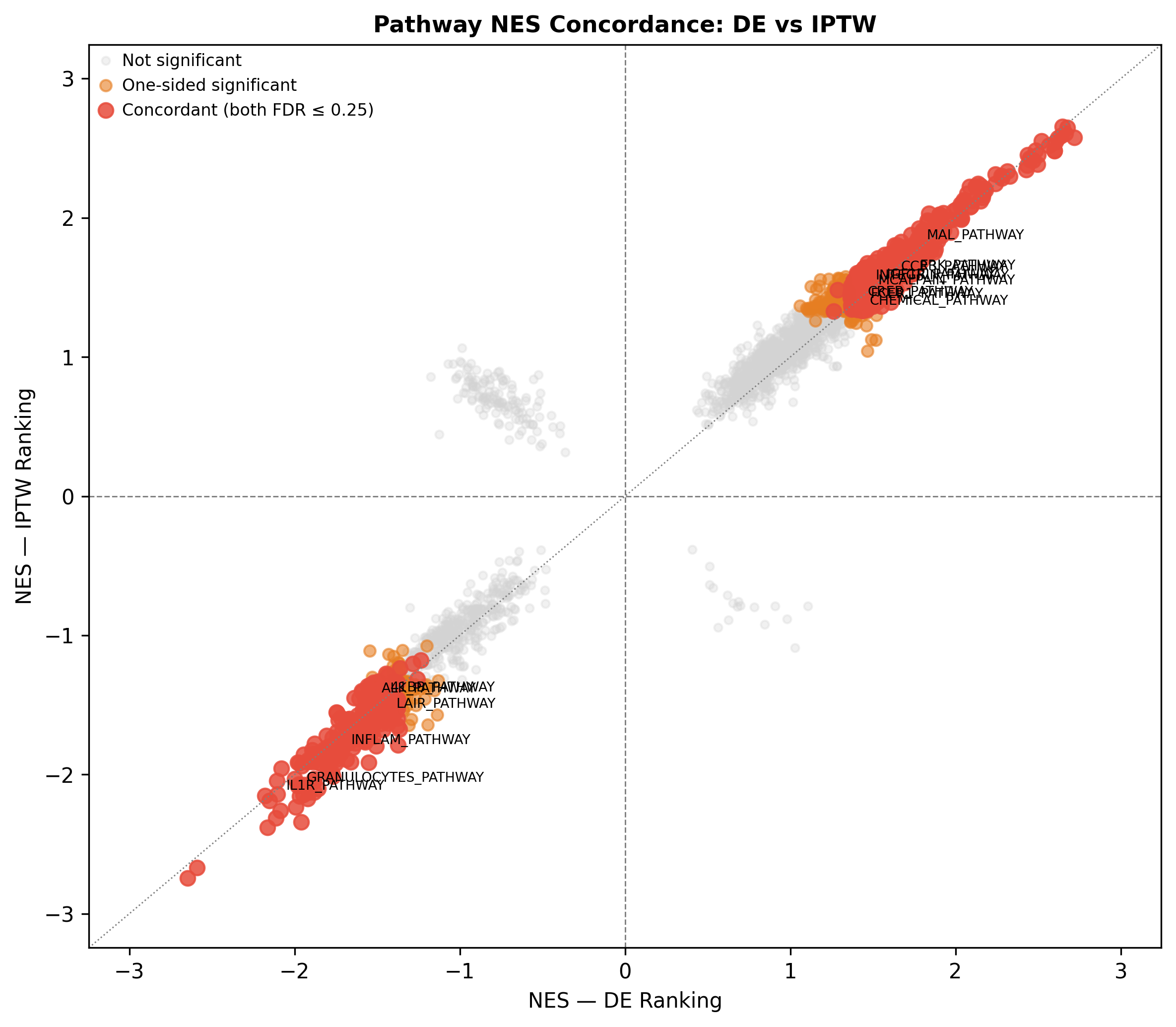
When the gene-level signal is confounding-bound, coordinated multi-gene programmes are not. Pathway-level enrichment, TF activity, and network modules converge on the same biology, recovering reproducible signal where single-gene FDR fails.
Two programmes, one mechanism axis
Programme A: Loss of colonocyte identity (lower in non-responders)
The WGCNA gainsboro module (n=2,026 genes; r_severity = −0.313, FDR 1.65×10⁻⁴) contains all the major epithelial hubs of the colonic mucosa:
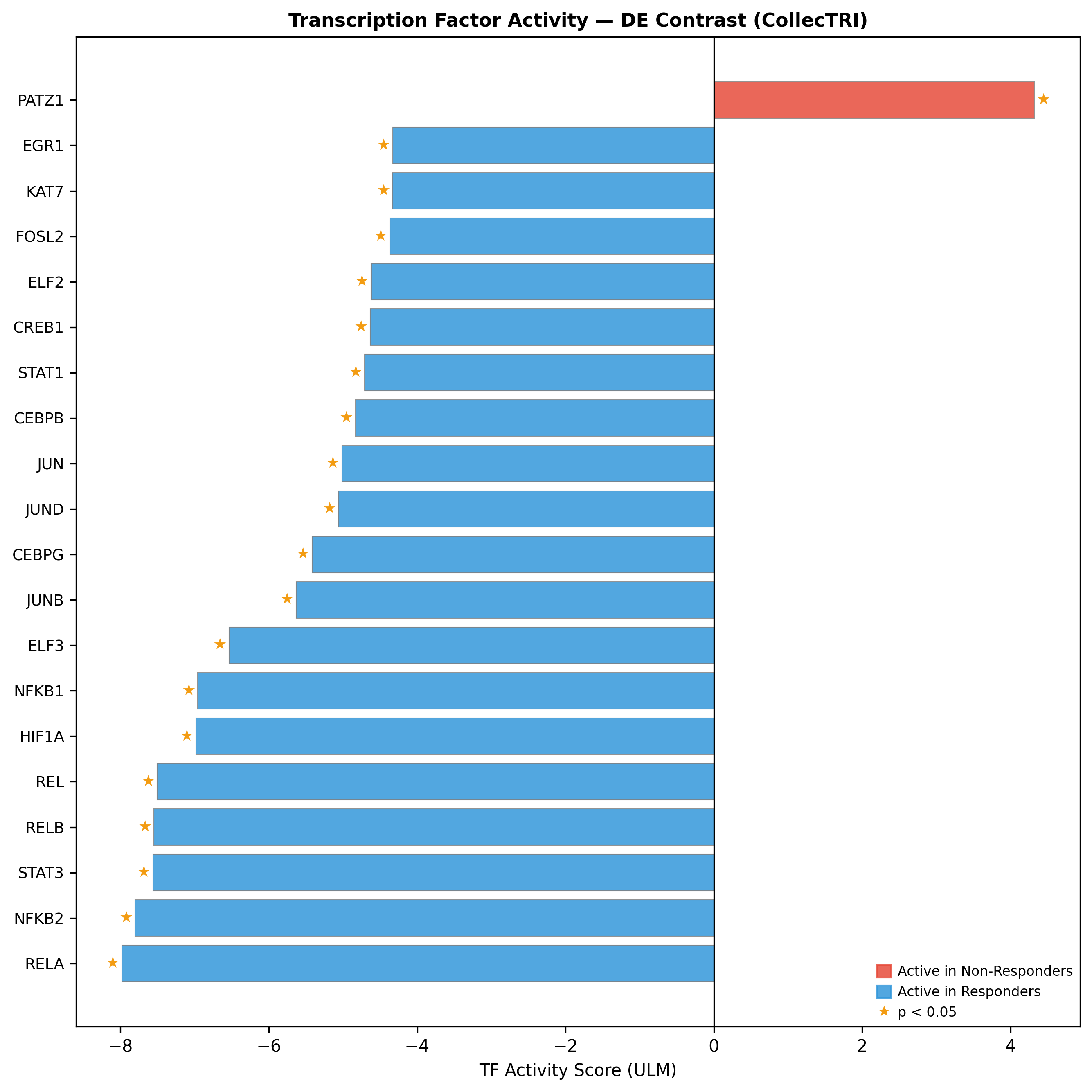
CDX2 is the #2-ranked responder-active TF (activity +4.23, p = 7×10⁻⁴), with CDX1, HNF1A, and FXR (NR1H4) co-activated. HALLMARK_OXPHOS NES +2.50 / +2.45; ribosome/translation programmes NES +2.66–2.72.
Programme B: Active inflammatory signalling (higher in non-responders)
HALLMARK_TNFA_SIGNALING_VIA_NFKB NES −2.59 / −2.67; IL6_JAK_STAT3 −1.54 / −1.71; complement cascade −2.11 / −2.31. The TF activity profile concentrates the inflammatory drivers tightly around four nodes:
| TF | Activity | p-value |
|---|---|---|
| RELA | -7.98 | <1e-12 |
| STAT3 | -7.56 | n/a |
| NFKB1 | -6.96 | n/a |
| HIF1A | -6.98 | n/a |
CXCL13 is the lead gene in 9 concordant non-responder-enriched pathways, with E-value 2.12 (ROBUST) across 3 independent evidence layers. M1 macrophage modules are higher in non-responders (Cohen's d = −0.28).
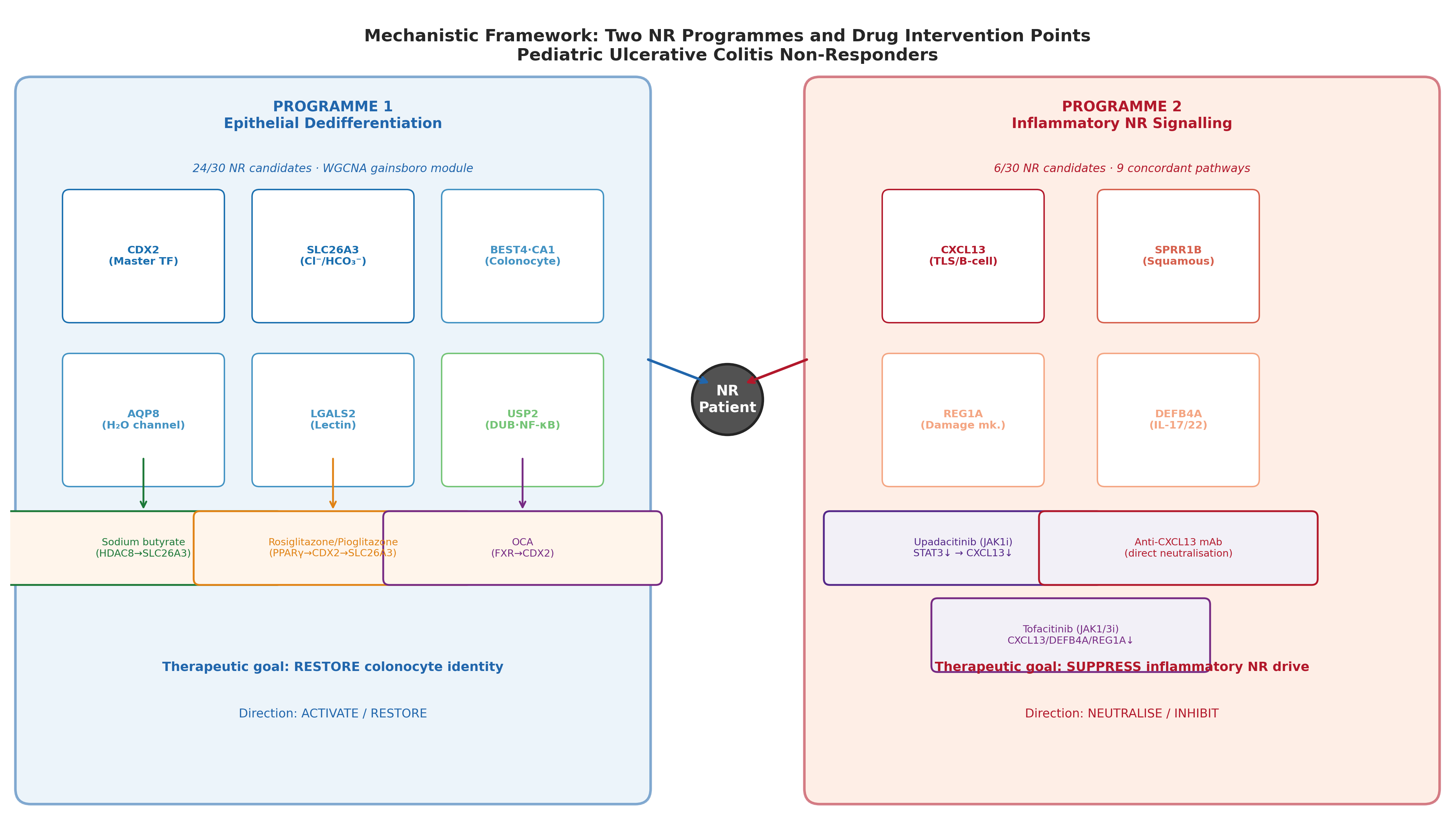
Step 4: Mechanistic convergence on the PPARγ → CDX2 → SLC26A3 axis
Four pharmacologically independent mechanisms all upregulate SLC26A3 / colonocyte identity. This kind of convergence is the strongest non-clinical prior we can give a candidate:
| Mechanism | Drug | Evidence | Reference |
|---|---|---|---|
| PPARγ agonism | Rosiglitazone | Phase 2 RCT n=105: response 44% vs 23% (p=0.04); remission 17% vs 2% (p=0.01) | PMID 18325386 |
| PPARγ agonism (topical) | Rosiglitazone enema | Mayo 8.9 → 4.3, restored mucosal PPARγ activity in vivo | PMID 20087330 |
| HDAC8 inhibition | Sodium butyrate | DSS rescue with Slc26a3 + tight-junction upregulation via HDAC8 → NF-κB | PMID 39440960 |
| Glucocorticoid receptor | Dexamethasone | Direct GRE in DRA promoter; rescues anti-IL-10R colitis | PMID 39657154 |
| FXR agonism | Obeticholic acid | Full DSS+TNBS protection in WT, not in Fxr-null mice | PMID 21242261 |
Restoring colonocyte identity via the PPARγ → CDX2 → SLC26A3 axis is reachable from at least four independent chemistry start points. Programme failure on safety at one node (e.g., rosiglitazone CHF) does not collapse the hypothesis.
Directionality, tractability, and pediatric translation
Step 3: The directionality constraint that breaks repurposing inventories
Of the 30 composite-prioritised candidates from network integration, 24 are higher in responders, meaning they need to be restored or activated in non-responders, not inhibited. This single observation disqualifies most off-the-shelf chemistry:
- FGFR3: T1-tractable (erdafitinib, infigratinib, pemigatinib, futibatinib) but counter-mechanistic for UC. Doubly disqualified for pediatrics: TYRA-300 selective FGFR3 inhibition lengthens long-bones in mice (Starrett 2025). Pediatric growth-plate contraindication is unmovable.
- CA1: All approved CAIs (acetazolamide, dorzolamide…) are inhibitors. Counter-mechanistic. Useful only as a pharmacodynamic biomarker.
- SLC26A3: Best ChEMBL IC50 25 nM, but only inhibitor series. Needs an activator/potentiator (CFTR-modulator analogy: ivacaftor for the related SLC26 superfamily).
This is where the analysis stops being a target list and starts being a chemistry strategy.
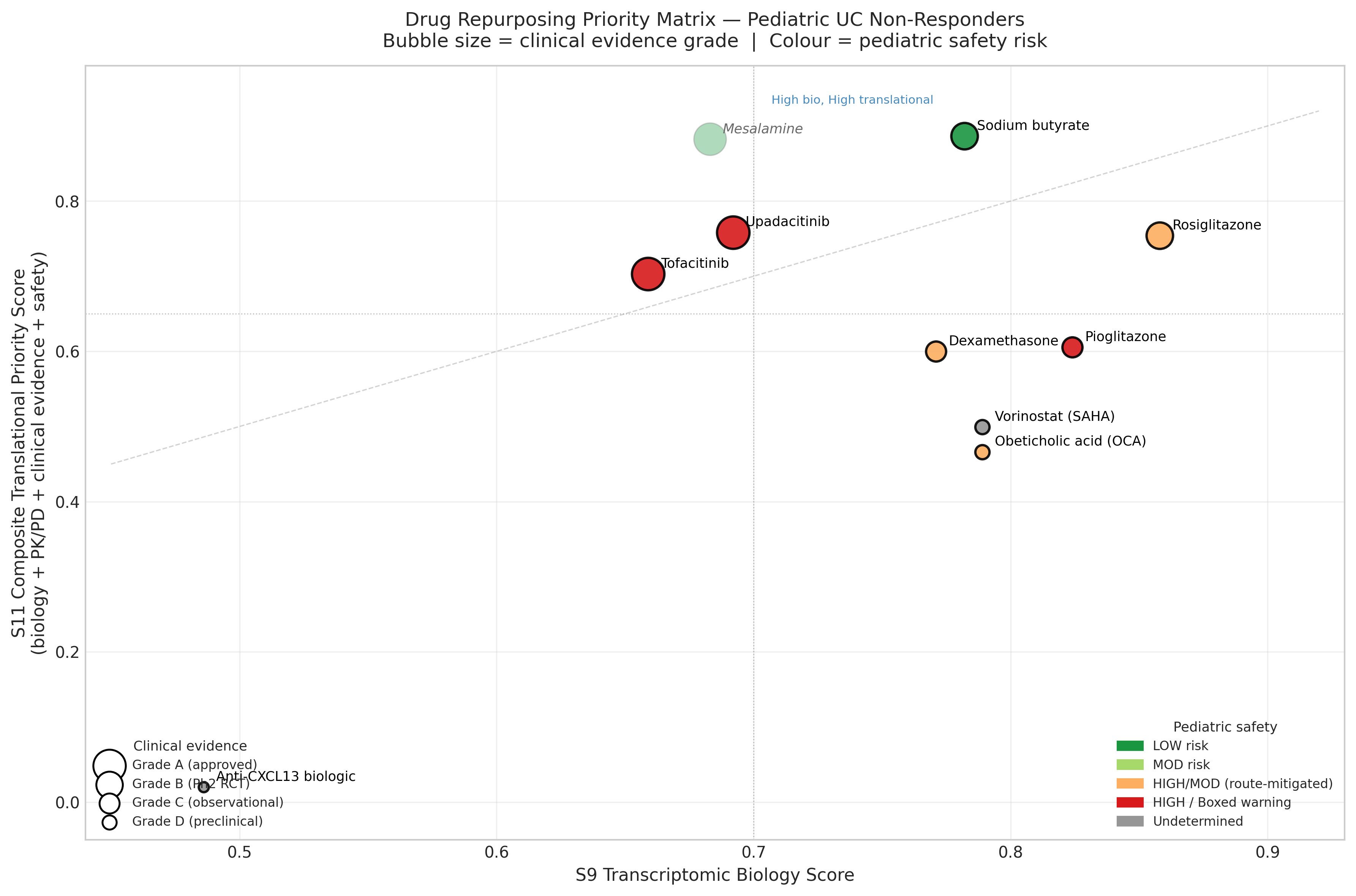
Step 5: Pediatric translational re-ranking inverts the biology-only order
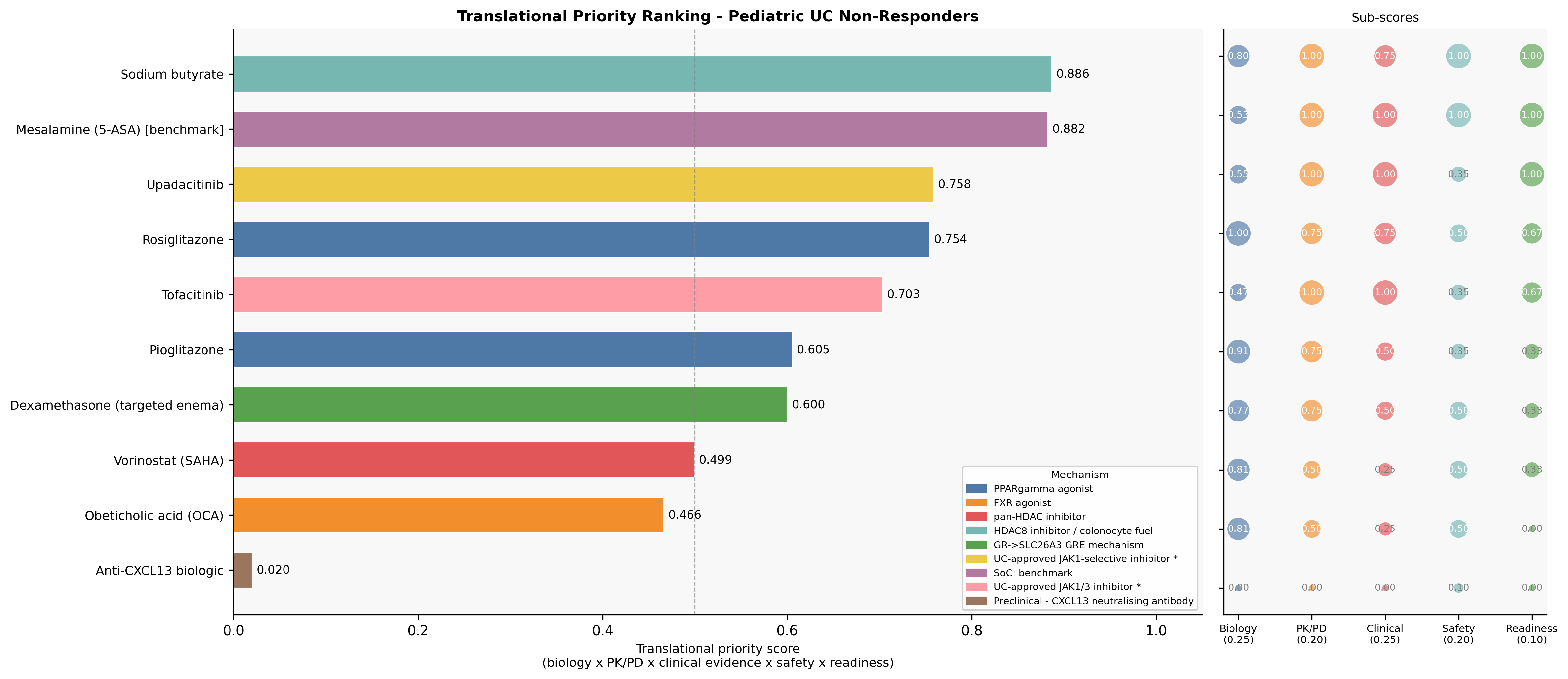
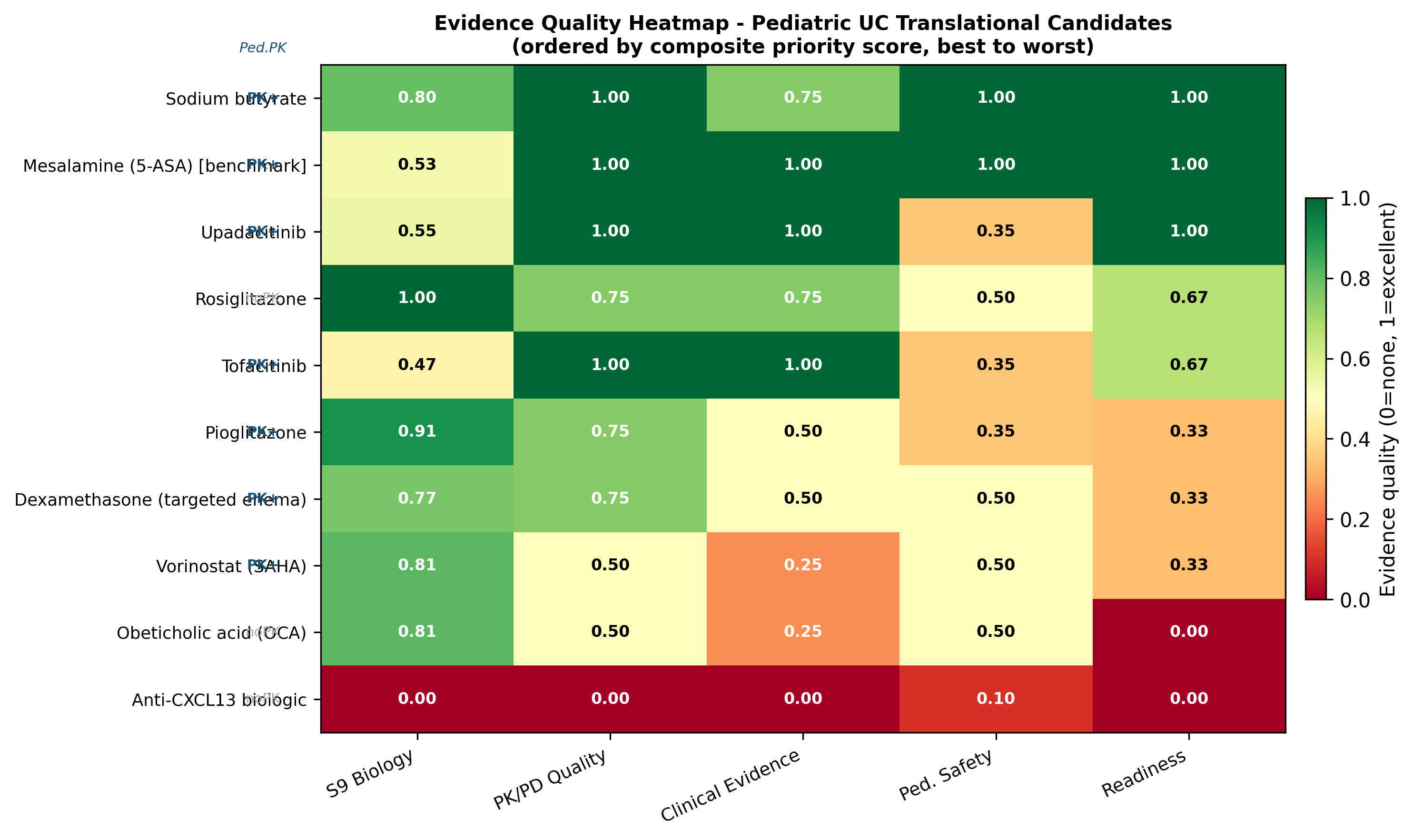
Biology is necessary but not sufficient. We re-scored the shortlist on a composite of biology (0.25), pediatric PK/PD (0.20), clinical evidence (0.25), pediatric safety (0.20), and translational readiness (0.10):
| Rank | Drug | Tier | S9 (biology) | S11 (translational) | Why the move |
|---|---|---|---|---|---|
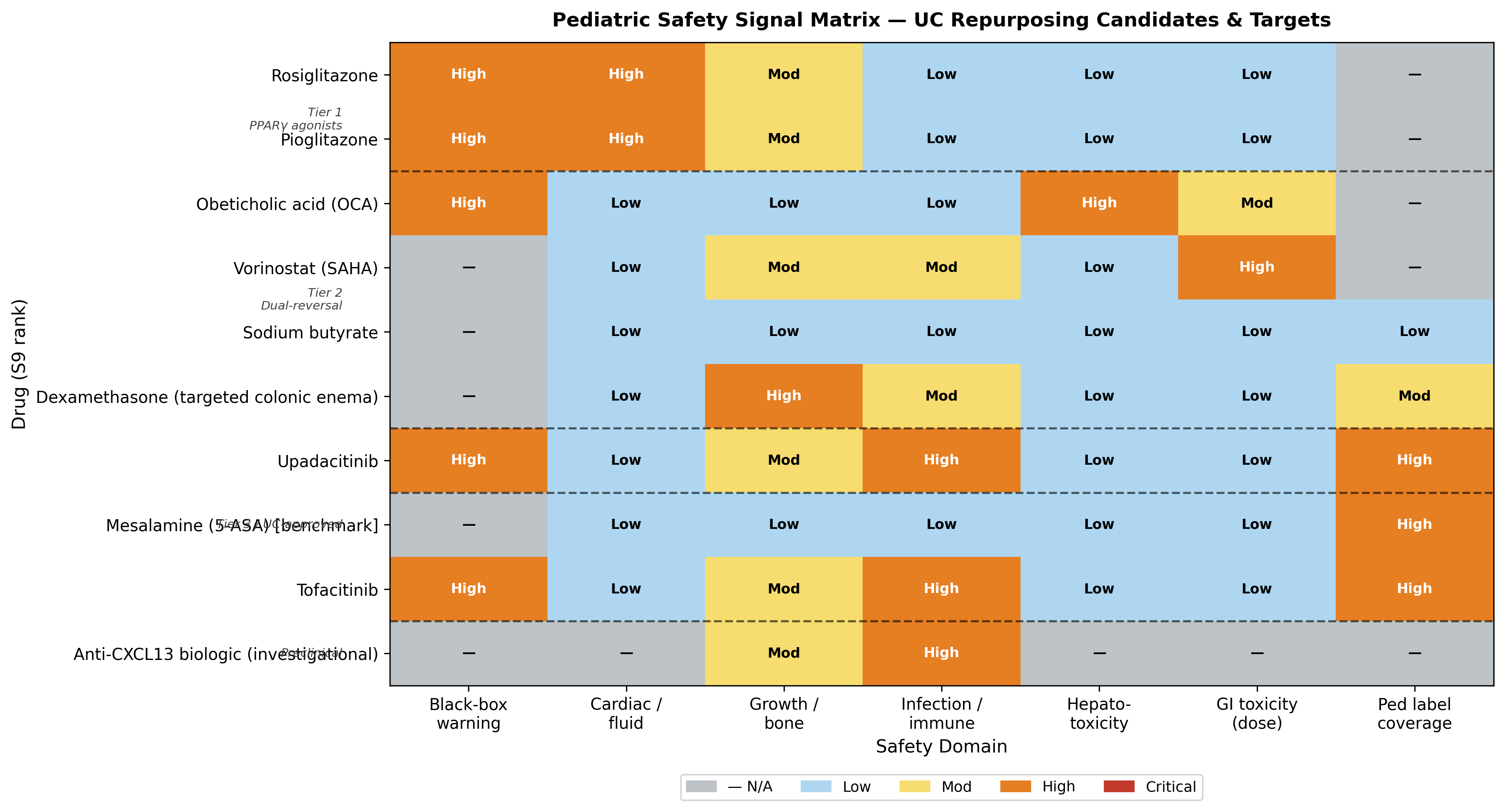
| 1 | Sodium butyrate (colonic-targeted enema or microencapsulated) | A | 0.782 | 0.886 | Only candidate with all-LOW pediatric safety domains; multi-mechanism convergence on SLC26A3 |
| 2 | Mesalamine (5-ASA) — benchmark | — | 0.683 | 0.882 | Standard of care comparator |
| 3 | Upadacitinib | B | 0.692 | 0.758 | Only pediatric-PK-validated JAK inhibitor; SELECT-YOUTH adult-equivalent AUC/Cmax; 84% response, 56% steroid-free remission at week 8 (n=100) |
| 4 | Rosiglitazone (enema) | B | 0.858 | 0.754 | Strongest biology, but no pediatric IBD PK + CHF black box demote it |
| 5 | Tofacitinib | C | 0.659 | 0.703 | Broader JAK1/3; preferred only if upadacitinib fails |
| 6 | Pioglitazone | C | 0.824 | 0.605 | Bladder-cancer signal disqualifying for chronic pediatric use |
Step 6: CXCL13, the highest-conviction novel target for biopharma drug discovery
If the question is where to invest dedicated discovery dollars, CXCL13 is the answer. It is the most ROBUST UP_NR driver in the entire analysis (E-value 2.12), lead gene in 9 concordant non-responder-enriched pathways, independently validated as an anti-TNF non-response marker in adult IBD biopsies (Iacucci 2023), and causal in chronic colitis-fibrosis: macrophage-derived CXCL13 drives fibrosis; shRNA knockdown rescues (Xiao 2025). No clinical-stage anti-CXCL13 agent exists for IBD.
The translational gate is real: CXCL13 organises Peyer's patches and germinal-centre B-cell responses, so chronic neutralisation in children requires dedicated developmental immunotoxicology (B-cell follicle ontogeny, mucosal IgA maturation). This is a launch-in-adults, study-down-to-pediatrics programme, but the target rationale is unusually strong for a chemokine.
Step 7: What gets disqualified, and why that matters
| Asset class | Verdict | Reason | Reference |
|---|---|---|---|
| FGFR3 inhibitors (erdafitinib et al.) | CRITICAL — disqualified | Counter-mechanistic (FGFR3 lost in NR) AND growth-plate contraindicated (TYRA-300 lengthens long-bones in WT mice) | PMID 40178985 |
| CA1 / pan-CA inhibitors | Counter-mechanistic | Use as pharmacodynamic biomarker only | n/a |
| Pioglitazone (chronic pediatric) | Demoted | Bladder-cancer signal incompatible with pediatric treatment horizon | n/a |
| Oral systemic butyrate at 150 mg bd | Refuted (formulation) | Neutral RCT in pediatric IBD; preserves the colonic-targeted hypothesis | PMID 36014789 |
The directionality framework is itself a methodological contribution: it filters repurposing inventories before mechanistic and safety review, removing 80% of the apparent T1 tractability for free.
What this means for biopharma decision-makers
This case study is, at heart, a discipline of constraints: causal-inference confounding bounds; directionality bounds chemistry; pediatric PK and growth-plate biology bound translation. Every constraint we honour eliminates a category of false starts.
Three programmes a biopharma portfolio could plausibly stand up tomorrow
Programme 1 — Colonic-targeted butyrate
Life-cycle / repositioning
- • Asset: pH-triggered microencapsulated or enema formulation
- • Pediatric study: PK / mucosal SCFA exposure & SLC26A3 mRNA induction in pediatric UC, age ≥6
- • Biomarker: mucosal SLC26A3, BEST4, AQP8, CA1 by RNA in situ — all part of Programme A
- • Risk: low; strategic value: foothold in pediatric UC with multi-mechanism rationale
Programme 2 — Pediatric upadacitinib for biologic-refractory UC
Existing asset, new positioning
- • Anchor: NCT05782907; SELECT-YOUTH PK validated; real-world data already supportive
- • Differentiator: targets Programme B specifically (CXCL13, REG1A, DEFB4A)
- • Mitigation: JAK1 selectivity reduces VZV / NK-depletion liability vs tofacitinib
Programme 3 — Anti-CXCL13 (or CXCR5 antagonist) discovery
Novel target, dedicated investment
- • Modality: neutralising mAb (preferred) or oral CXCR5 antagonist
- • Adult-first strategy: biologic-refractory IBD with elevated baseline mucosal CXCL13 as enrichment biomarker
- • Pediatric path: dedicated developmental immunotoxicology before pediatric IND
- • Differentiator: only highest-confidence ROBUST inflammatory driver with no clinical-stage competitor in IBD
Stretch programmes
- SLC26A3 potentiator / activator HTS — CFTR-modulator analogy; tractable pocket confirmed by 25 nM inhibitor series; needs orthogonal activator screen.
- Topical / colonic-restricted PPARγ agonist — bypasses the rosiglitazone CV liability; pediatric IBD PK study is the gating experiment.
Limitations
- Single dataset (PROTECT GSE109142, n=206); no independent pediatric UC replication cohort.
- Therapy × severity collinearity (χ² = 139.88) is structurally irreducible; no candidate reaches ROBUST_STRONG (E ≥ 3 with attenuation ≥ 0.75).
- Bulk RNA-seq cannot distinguish loss of the BEST4⁺ colonocyte subset from transcriptional suppression within it — single-cell follow-up required.
- No LINCS/CMap connectivity in this analysis; repurposing was target-centric.
- Composite scores are screening-level prioritisation, not regulatory benefit-risk.
- Week-4 readout; biology predicting durable (week-52) outcomes not separately modelled.
The translational-medicine takeaway
The PROTECT analysis would have failed the classical "find your top 10 FDR genes and run a target ID" workflow: there are no top 10 FDR genes. What it succeeded at was layering causal-inference adjustment, multi-modality enrichment, network integration, directionality-aware tractability, and pediatric PK/safety re-weighting to surface a shortlist of three actionable programmes and one disqualification with a compelling mechanistic explanation for each.
For a biopharma audience, the lesson generalises: when the cohort is confounded (and most disease cohorts are), the path forward isn't a bigger gene list, it's a denser evidence stack that respects directionality and child-specific physiology. The candidates that survive that filter are worth reaching for.