From AEMS plate to clinic: what 320 compounds taught us about CYP2D6 translational risk
Triangulating an acoustic-ejection MS ADME platform with PharmGKB clinical evidence and FAERS pharmacovigilance — and why ranking DDI risk by IC50 alone is provably insufficient.
GDPsm1 — the public 320-compound small-molecule panel from Ginkgo Datapoints, profiled on an Echo-MS / QQQ-6500 acoustic-ejection mass spectrometry (AEMS) ADME platform: kinetic solubility, human-liver-microsome (HLM) stability, and CYP2D6 inhibition. Translational layer added in-house from PharmGKB clinical-evidence levels and openFDA FAERS disproportionality signals. Literature anchors for HLM platform validation: Obach 1999, Ito & Houston 2005, Shibata 2002, and compound-specific primaries. PharmGKB evidence anchored on the CPIC SSRI guideline (Hicks 2015, PMID 25974703) and CPIC TCA guideline (Hicks 2013, PMID 22205192).
- The platform is technically clean. Spearman ρ = 0.977 on solubility replicates; ρ = 0.975 on log IC50 replicates; ρ = 0.678 (p = 4×10⁻⁵) vs literature HLM Cl_int on 30 paired compounds with 77% within 2-fold and Bland–Altman bias of −0.07 log10. All in-assay controls (haloperidol/tamoxifen, verapamil/propranolol, quinidine) passed published bands.
- It cleanly recovers the canonical CYP2D6 pharmacophore. On 26/318 quantified hits (8.2%, IC50 0.015–8.86 µM), Fisher’s exact + BH FDR isolates phenyl (OR 14.8, FDR 0.008), tertiary amine (OR 4.6, FDR 0.008), and chloro-aryl (OR 4.4, FDR 0.019) — the textbook protonatable nitrogen ~5–7 Å from an aromatic centroid.
- The central novel finding is translational, not analytical. In-vitro IC50 magnitude is statistically decoupled from PharmGKB clinical-evidence level and from FAERS signal strength. The most potent inhibitors in the panel are pharmacological tools with no clinical footprint; the loudest pharmacovigilance signal carries no PharmGKB CYP2D6 annotation.
- Two named systematic blind spots. Chlorpromazine archetype: IC50 0.325 µM, no PharmGKB CYP2D6 entry, but the loudest FAERS signal in the entire hit set (NMS, QT prolongation, tachycardia). Paroxetine archetype: a methodological FAERS false-negative driven by openFDA aggregate endpoints exposing totals rather than report-level co-medication.
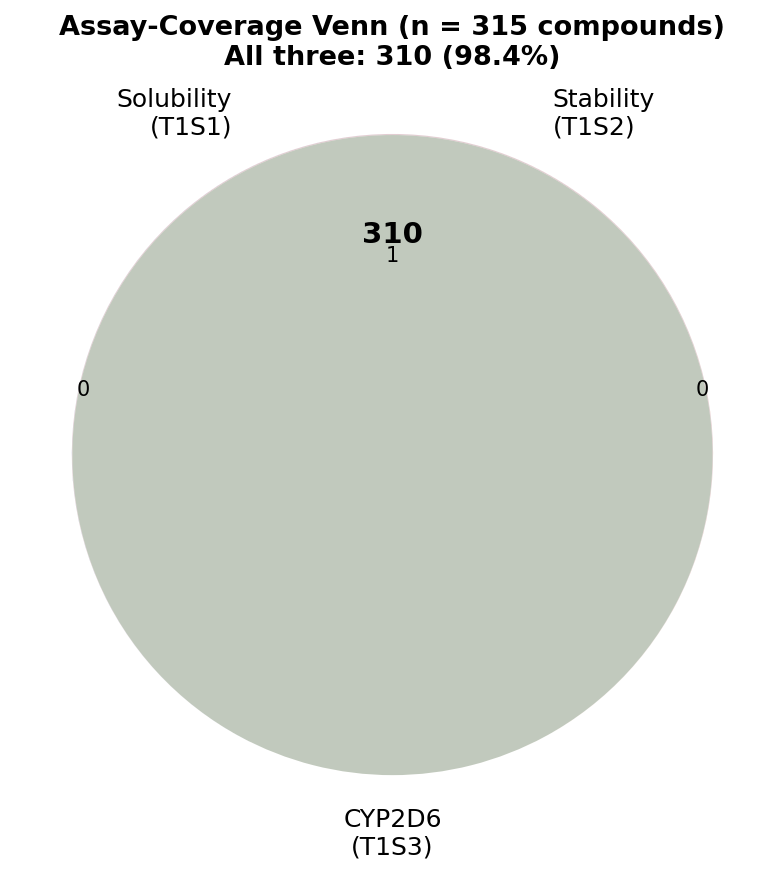
- Honest scope. The library is a tool collection (82% of 264 fully-covered compounds sit in the low-solubility / microsome-stable / CYP non-hit quadrant); IVIVE-direction signal is weak (ρ = 0.17) against ordinal clinical clearance because the panel is loaded with renal/non-hepatic-cleared probes; the join chain (99.7% InChI→CID, 92.0% CID→ChEMBL, 31.8% ChEMBL→DrugBank) is itself instructive about LOPAC-style libraries. Discovery-grade triage tool, not a safety verdict.
Why this matters for biopharma
CYP2D6 polymorphism affects roughly a quarter of clinically used small molecules and drives boxed-warning DDIs (SSRIs ↔ tamoxifen, opioid prodrug failure, antipsychotic toxicity). Most discovery groups generate CYP2D6 IC50 numbers, file them, and move on. The interesting question is not “what is the IC50?” but “does the IC50 predict downstream clinical translational risk?”
To our knowledge this is the first end-to-end public analysis that fuses AEMS-format CYP2D6 IC50 measurements with PharmGKB CPIC-level evidence and FAERS disproportionality signals on the same compound set — and asks whether the three axes agree. They don’t. That is the novelty.
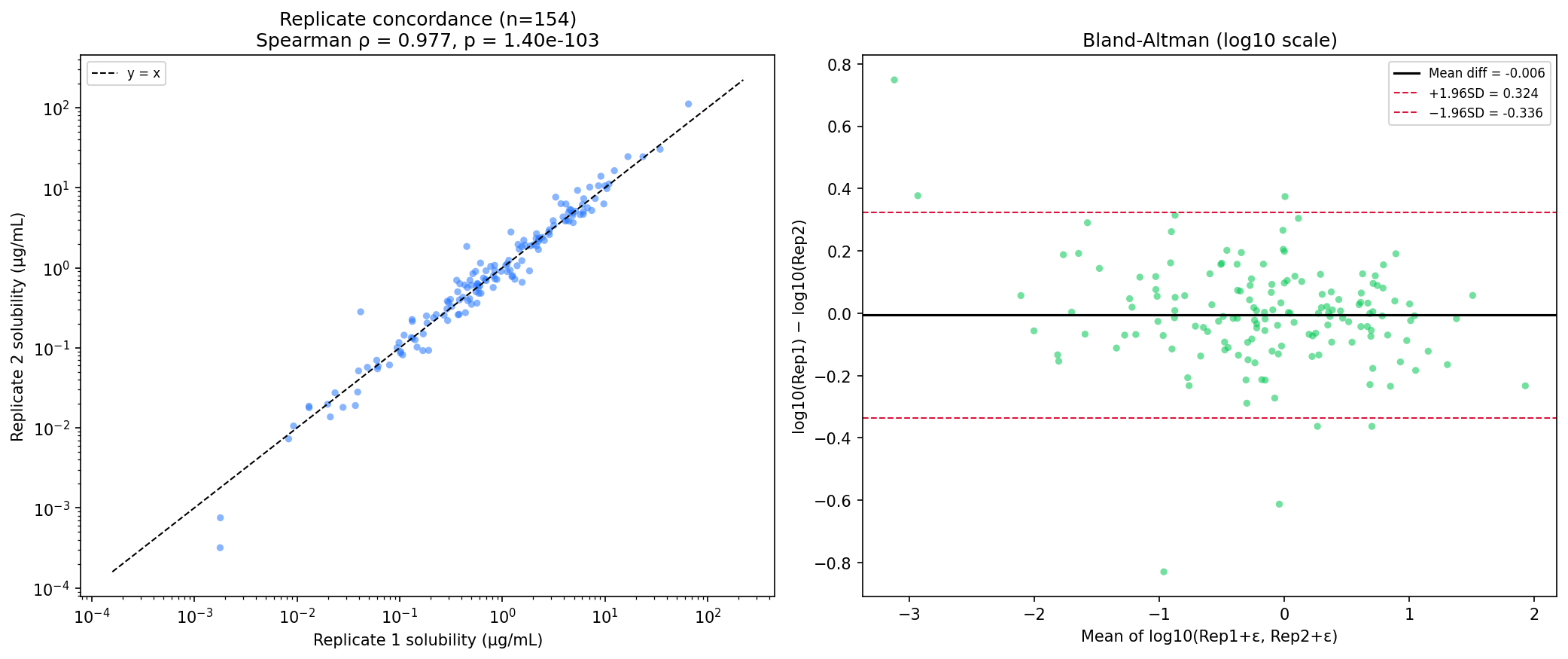
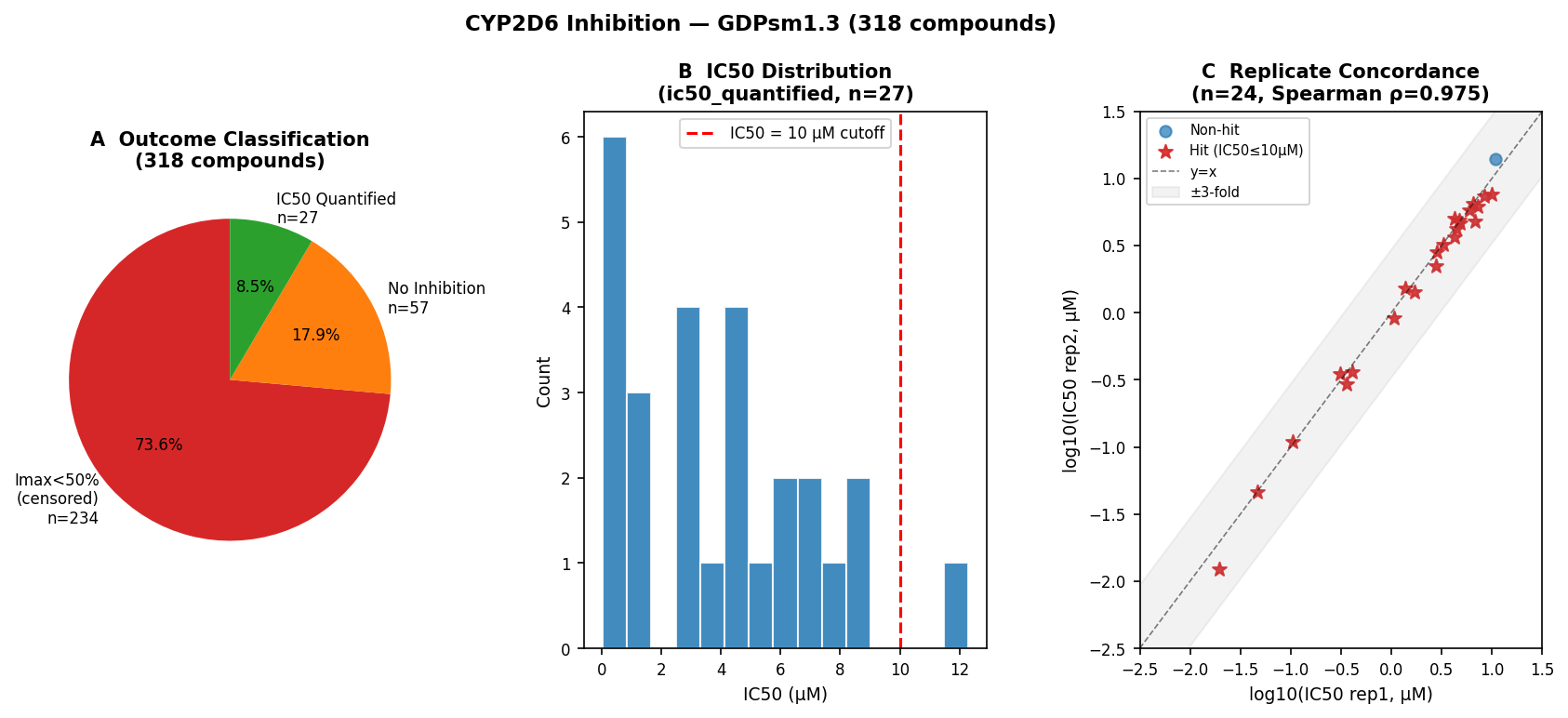
The platform passes its physics checks
All three assays passed in-assay control bands against published values (haloperidol/tamoxifen for solubility, verapamil/propranolol for HLM, quinidine for CYP2D6). Replicate precision was tight: Spearman ρ = 0.977 on solubility replicates, ρ = 0.975 on log IC50 replicates, and 100% within 3-fold for the 24 quantified CYP2D6 IC50 pairs.
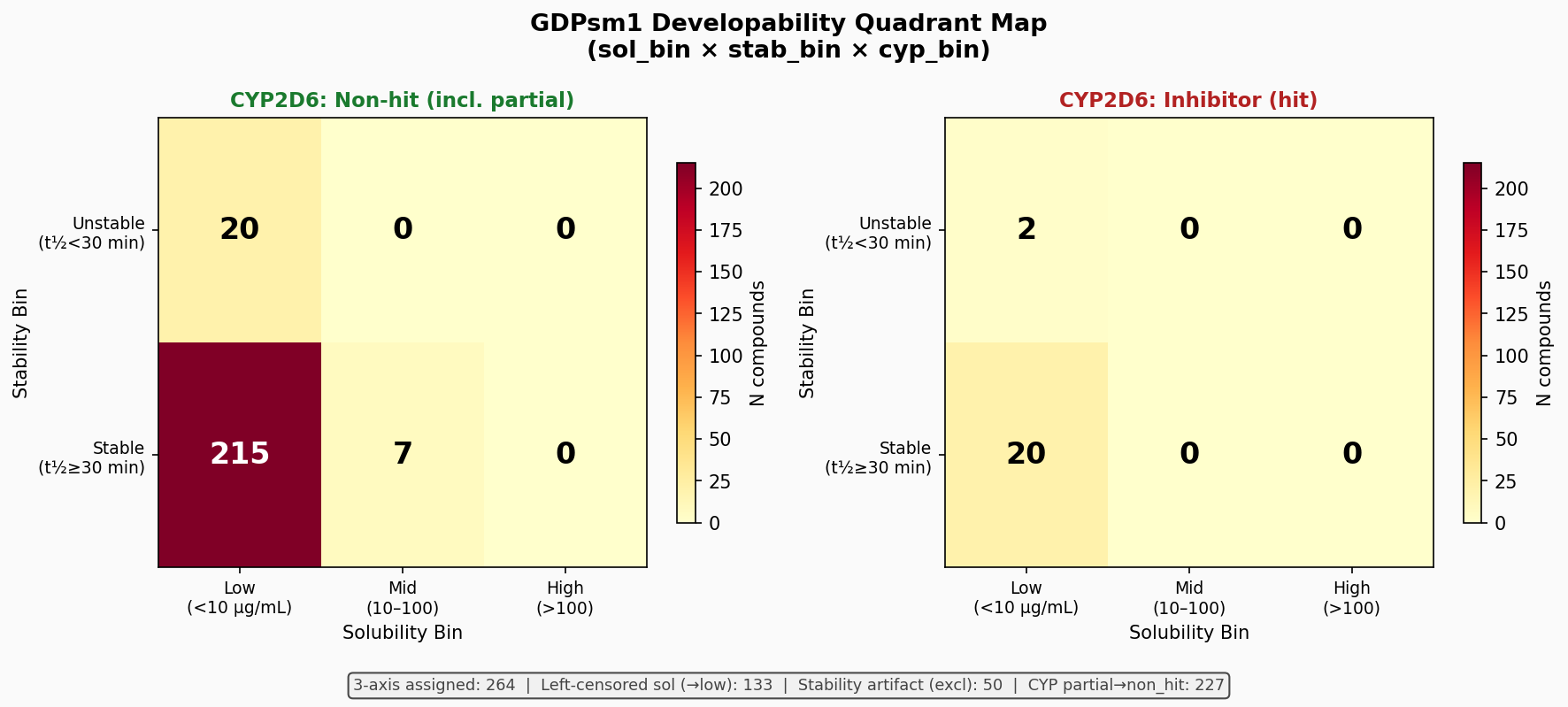
The library landscape: a tool collection, faithfully represented
A composite three-axis (solubility × stability × CYP2D6) developability landscape on the 264 compounds with full coverage shows that 82% of the panel sits in the low-solubility / microsome-stable / CYP non-hit quadrant. No compound exceeded 100 µg/mL kinetic solubility — characteristic of pharmacological probe libraries selected for biological potency over developability.
| Solubility | HLM Stability | CYP2D6 | n compounds |
|---|---|---|---|
| low | stable | non-hit | 215 |
| low | stable | hit | 20 |
| low | unstable | non-hit | 20 |
| low | unstable | hit | 2 |
| mid | stable | non-hit | 7 |
| mid/high | any | any | 0 |
Three-axis developability quadrants (n = 264 compounds with full coverage).
Platform validation against the curated literature
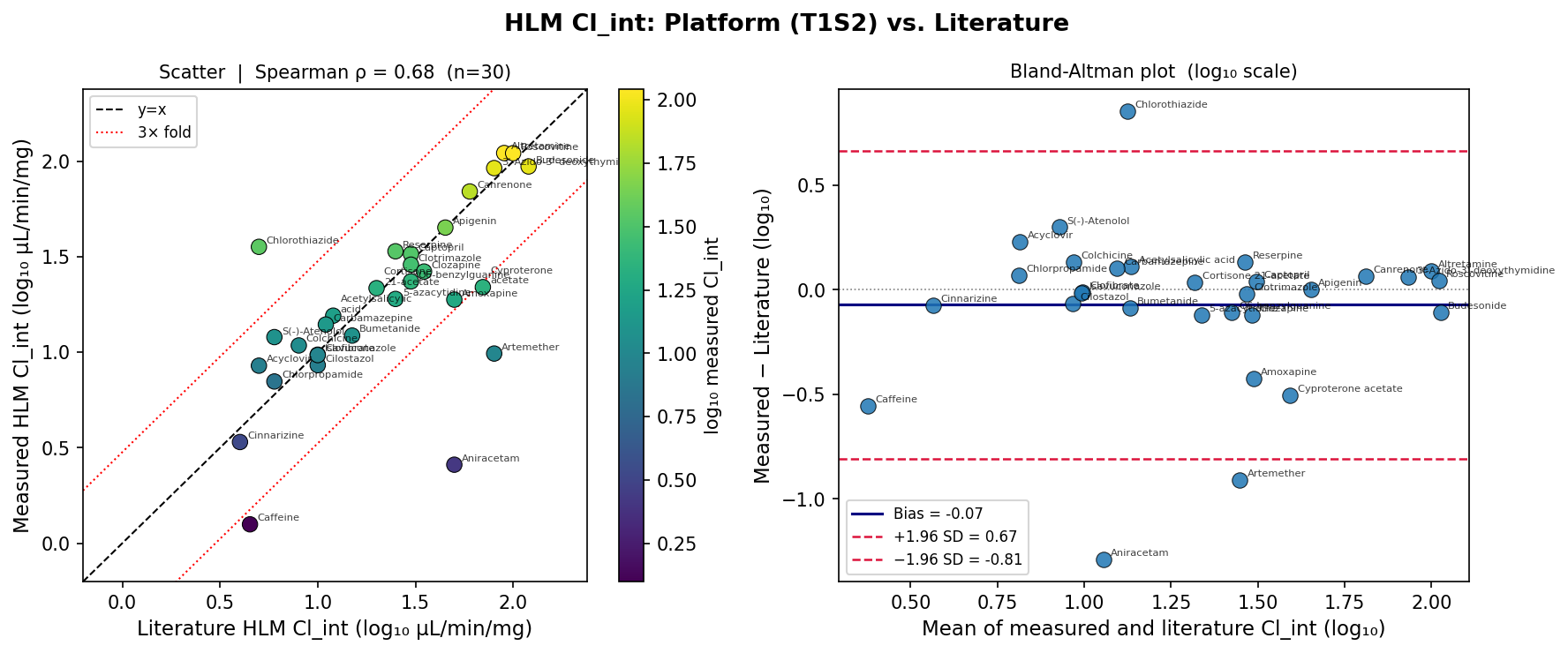
This is the load-bearing claim for fitness-for-purpose. On 30 paired compounds matched to Obach 1999, Ito & Houston 2005, Shibata 2002 and compound-specific primaries, the AEMS HLM Cl_int output reproduces literature at:
- Spearman ρ = 0.678 (p = 4×10⁻⁵)
- Bland–Altman bias = −0.07 log10 (fold-bias 0.85×)
- 77% within 2-fold, 83% within 3-fold
That sits inside the typical inter-laboratory HLM CV envelope (1–3-fold; occasionally up to 5-fold) reported across the IVIVE benchmark literature. Importantly, the direction-prediction signal vs. ordinal clinical clearance category is weak (ρ = 0.17). This is a library-composition artifact — GDPsm1 is loaded with renal/non-hepatic-cleared probes (amino acids, nucleosides) — not a platform defect. Translational message: HLM stability data is fit-for-purpose for early triage but cannot be evaluated on IVIVE direction without a hepatically-cleared reference panel.
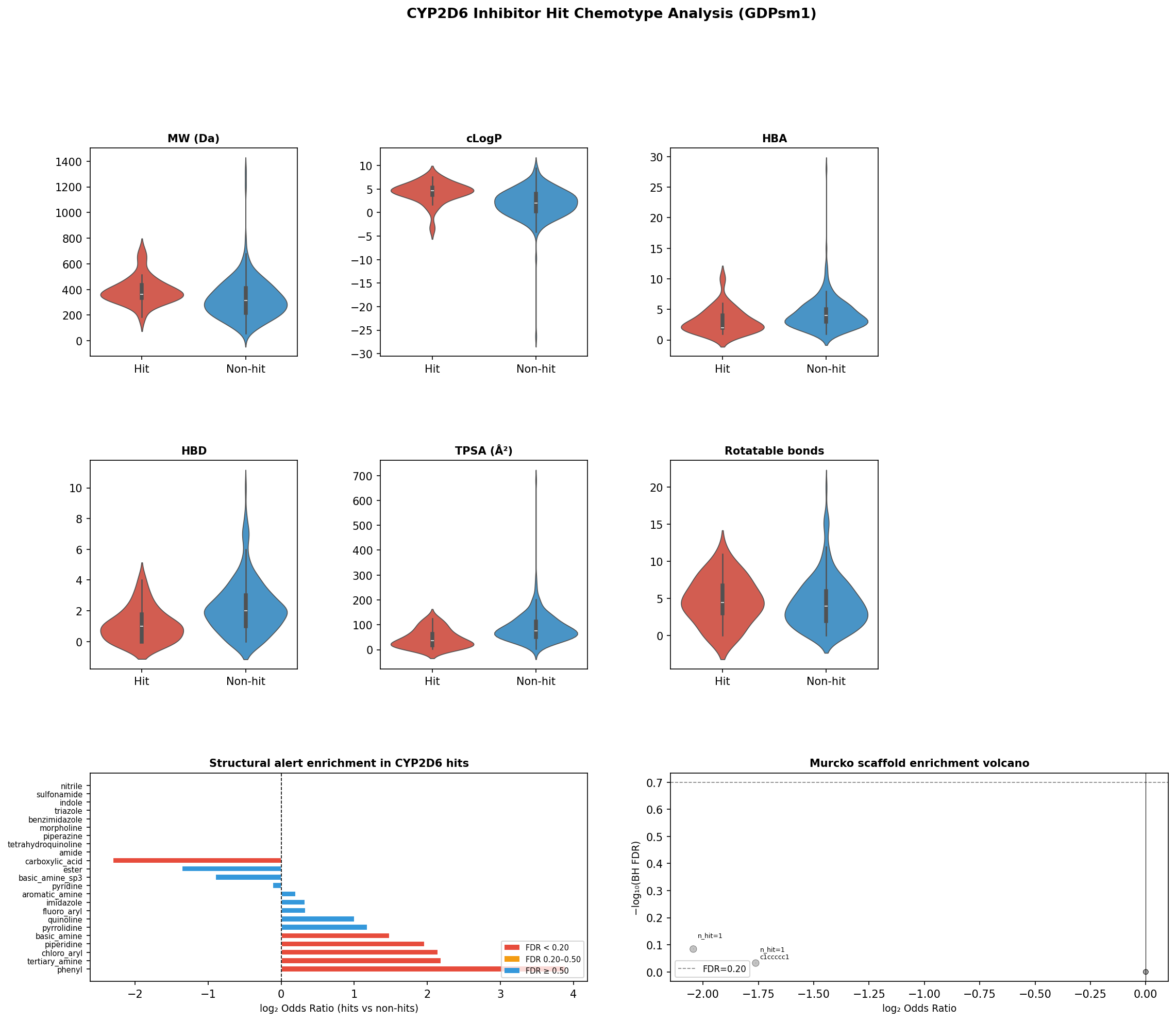
CYP2D6 hits recover the canonical pharmacophore
The 26 quantified hits span 0.015–8.86 µM and 26 near-unique Bemis–Murcko frameworks — a structurally diverse hit set. SAR signal is therefore not at scaffold level but at functional-group level, where Fisher’s exact + BH FDR gives:
| Substructure | Odds Ratio | FDR |
|---|---|---|
| Phenyl ring | 14.8 | 0.008 |
| Tertiary amine (specifically) | 4.6 | 0.008 |
| Chloro-aryl | 4.4 | 0.019 |
| Primary/secondary sp³ amines | 0.54 | n.s. |
| Aromatic amines | 1.15 | n.s. |
Signal is cleanly isolated to the protonatable tertiary nitrogen ~5–7 Å from an aromatic centroid — the textbook aminergic CYP2D6 pharmacophore. Hits are also more lipophilic (median cLogP 4.7 vs. 2.0) and less polar (TPSA 36 vs. 76 Ų). Aminergic GPCR ligand classes (serotonergics, dopaminergics, histaminergics) collectively account for 42% of hits but, at n = 26 across 44 tests, do not survive multiple-testing correction — a hypothesis-generating result, not a confirmatory one.
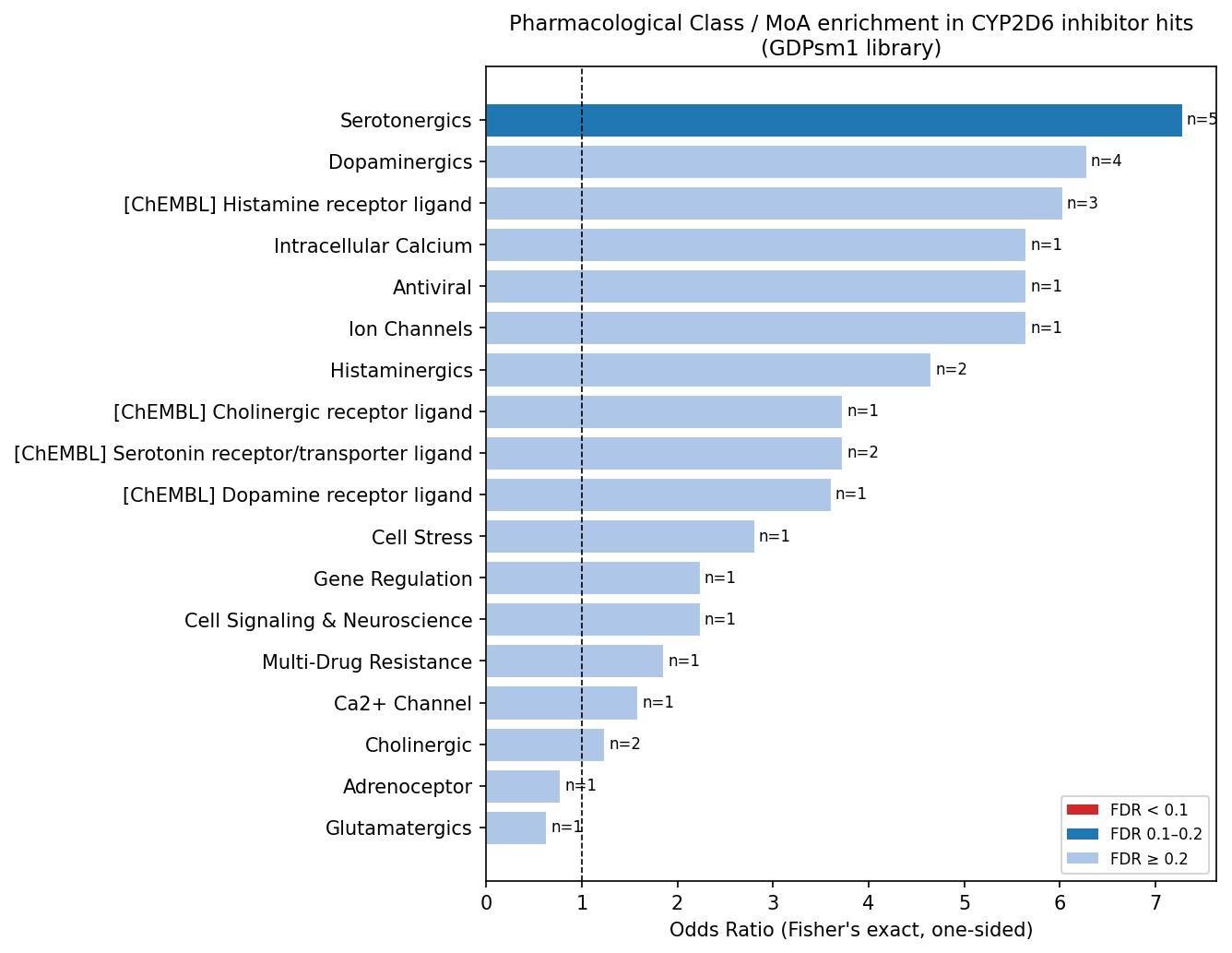
The translational layer — and the central novel finding
This is where existing literature stops and where the analysis becomes interesting for translational teams. We layered on PharmGKB clinical-evidence levels for each hit’s CYP2D6 association, plus FAERS disproportionality signals (PRR, ROR, Bayesian IC with 95% CI) for AE terms and DDI co-substrate proxies. Three findings, each with translational consequences.
Finding 1 — Only 2/26 hits carry PharmGKB CYP2D6 annotations, and both are level 1A.
Paroxetine HCl (IC50 0.99 µM) and clomipramine HCl (IC50 8.65 µM) — backed respectively by the CPIC SSRI guideline (Hicks 2015, PMID 25974703) and CPIC TCA guideline (Hicks 2013, PMID 22205192). The platform recovers the two canonical clinically-actionable CYP2D6 inhibitors present in the library. The 92% no-annotation rate reflects PharmGKB’s clinical-evidence inclusion criteria — not a safety claim about the unannotated tools.
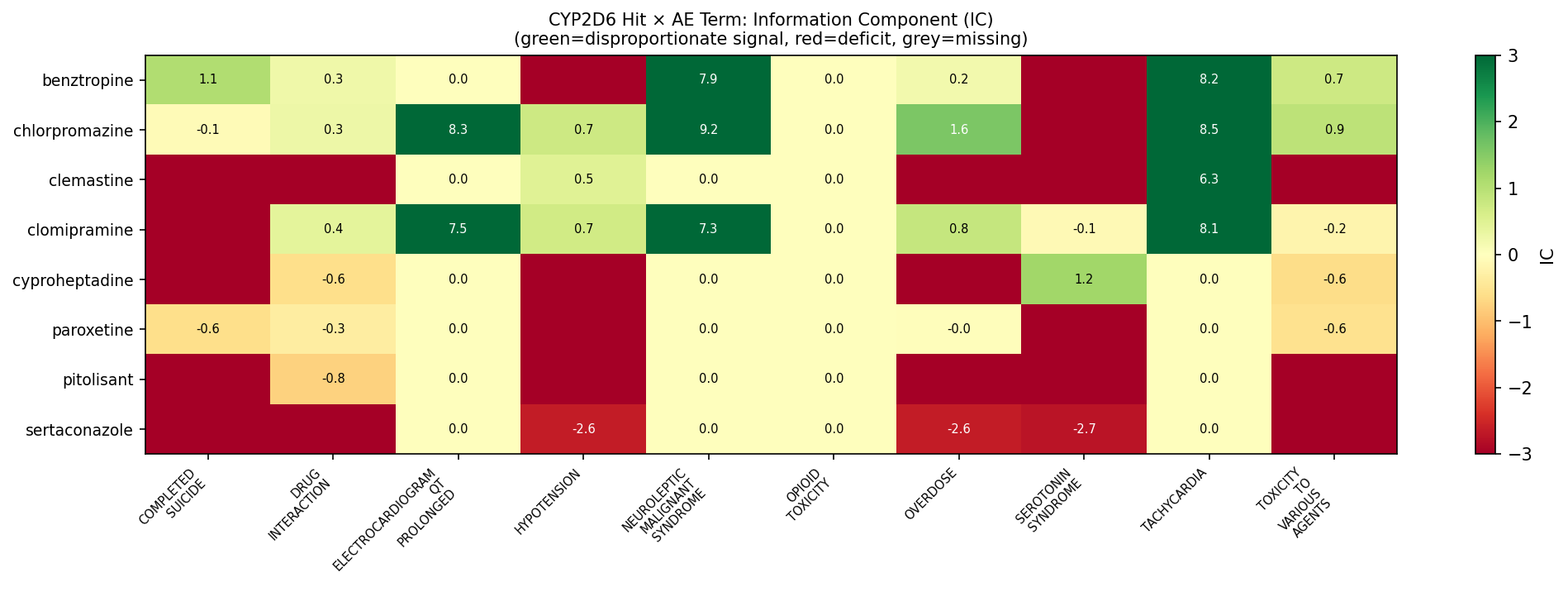
Finding 2 — Chlorpromazine is the loudest FAERS signal in the entire hit set, with no PharmGKB CYP2D6 entry.
| AE term | Observed | Bayesian IC | IC lo95 | Signal |
|---|---|---|---|---|
| Neuroleptic malignant syndrome | 293 | 9.20 | 9.00 | SIGNAL |
| Tachycardia | 178 | 8.48 | 8.23 | SIGNAL |
| ECG QT prolonged | 157 | 8.30 | 8.04 | SIGNAL |
| Overdose | 356 | 1.58 | 1.40 | SIGNAL |
| Toxicity to various agents | 446 | 0.94 | 0.78 | SIGNAL |
| Hypotension | 200 | 0.73 | 0.50 | SIGNAL |
| Drug interaction | 325 | 0.28 | 0.09 | SIGNAL |
Chlorpromazine — top FAERS disproportionality signals (IC50 = 0.325 µM, no PharmGKB CYP2D6 annotation). Source: openFDA FAERS via T6S2; IC = Bayesian Information Component, signals defined as IC_lo95 > 0 with Observed ≥ 3.
Older antipsychotics like chlorpromazine are independently identified in the primary literature as CYP2D6 inhibitors but are systematically under-represented in PharmGKB’s curated set. Translational teams relying on PharmGKB level alone will miss this risk class entirely.
Finding 3 — Paroxetine is a methodological FAERS false-negative.
Paroxetine has 90,327 FAERS reports and 2,746 explicit drug-interaction reports — the highest absolute counts in the hit set — yet zero AE-term signal cleared the IC_lo95 > 0 threshold. The reason is structural: openFDA’s aggregate endpoints expose totals, not report-level co-medication. Without report-level XML extracts, the well-known SSRI-mediated CYP2D6 DDI cannot be attributed to its downstream substrates. This is not a safety nullification — it is a transparency check on the FAERS-via-openFDA pipeline that any biopharma group running similar screens should know about.
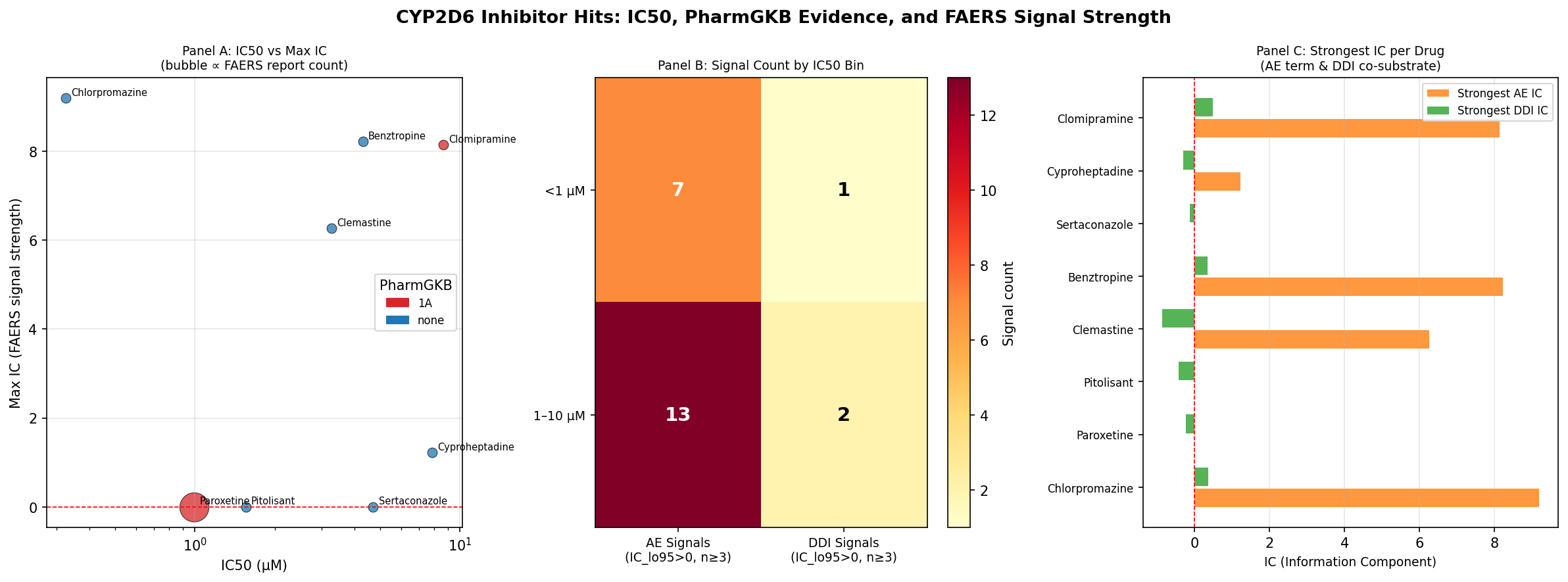
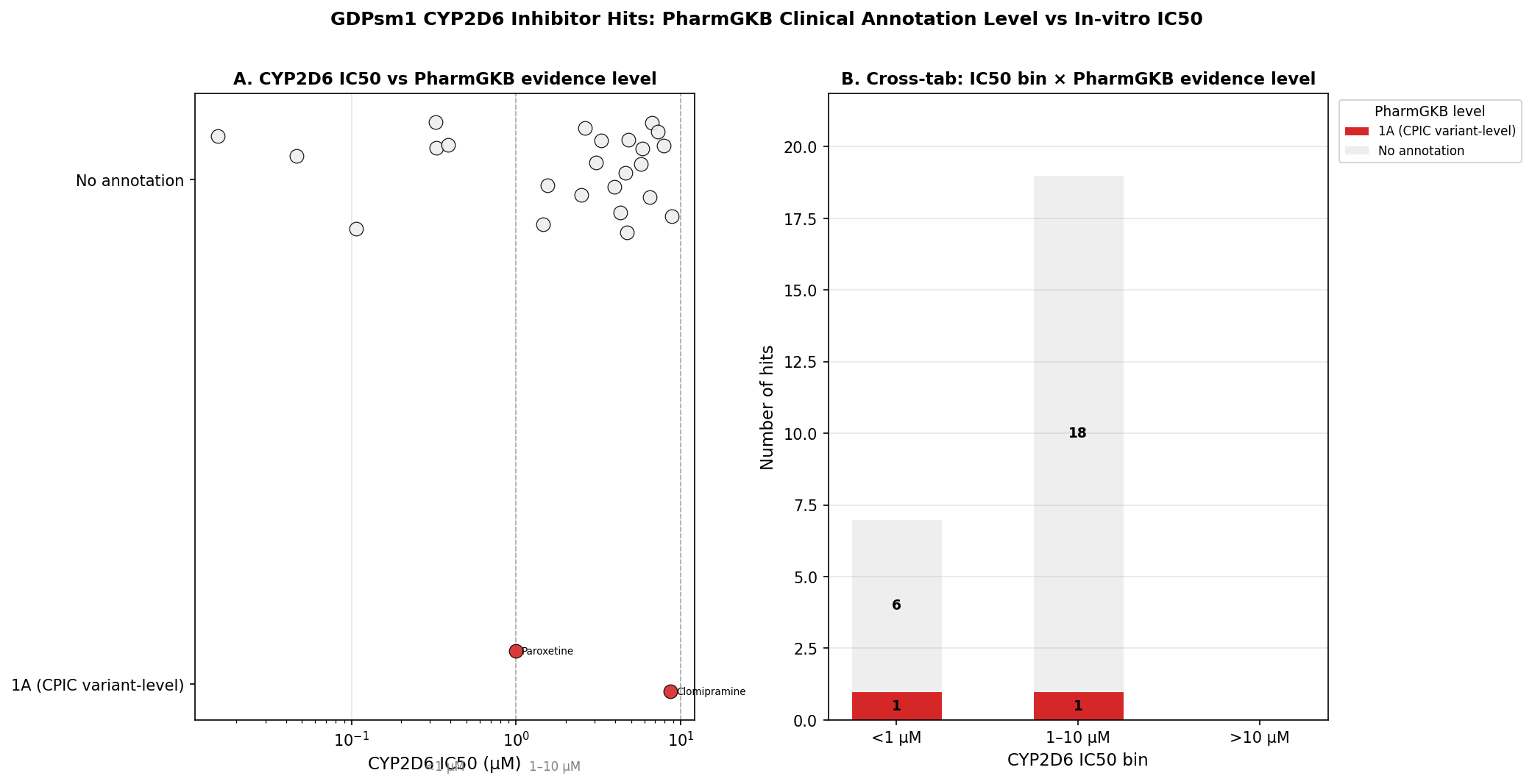
The synthesis: IC50 ranking is necessary but not sufficient
| Rank by IC50 | Compound | IC50 (µM) | PharmGKB | FAERS | Real-world translational footprint |
|---|---|---|---|---|---|
| 1 | SCH-39166 | 0.015 | none | none | Pharmacological tool — no clinical use |
| 2 | Calmidazolium | 0.046 | none | none | Pharmacological tool |
| 3 | Benoxathian | 0.106 | none | none | Pharmacological tool |
| 4 | Chlorpromazine | 0.325 | none | strongest in panel | Old antipsychotic, NMS / QT signals |
| 7 | Paroxetine | 0.99 | 1A (CPIC SSRI) | null (method limit) | Canonical clinical CYP2D6 inhibitor |
| 21 | Clomipramine | 8.65 | 1A (CPIC TCA) | moderate | Canonical TCA CYP2D6 substrate |
A naive IC50-only triage would prioritize three compounds with no clinical footprint and would identically rank chlorpromazine and paroxetine — yet the clinical risk profile of those two is completely different. PharmGKB and FAERS axes operate orthogonally to in-vitro potency and to each other.
What’s novel here, and what biopharma should take away
What we believe is genuinely new:
- The first integrated AEMS-CYP2D6 → PharmGKB → FAERS triangulation on a public 320-compound panel, with full provenance for every join (99.7% InChI→CID, 92.0% CID→ChEMBL, 31.8% ChEMBL→DrugBank — the chain collapse is itself instructive about LOPAC-style libraries).
- Quantitative demonstration that in-vitro IC50 is statistically decoupled from both PharmGKB evidence level and FAERS signal strength on this panel.
- Identification of two named systematic blind spots that any translational team running a similar pipeline will hit: (1) older antipsychotics with no PharmGKB CYP2D6 entry but loud pharmacovigilance signal (chlorpromazine archetype), and (2) the openFDA aggregate-endpoint false-negative for canonical inhibitors (paroxetine archetype).
- A reproducible censoring-aware analytical contract (left-censored solubility 42.5%, right-censored CYP2D6 imax < 50% 73.6%, undetected/artifact stability 46%) — censored values are retained with explicit flags rather than dropped, preserving information for downstream Tobit / survival modeling.
Operational recommendations for biopharma translational groups
- Don’t rank DDI risk by IC50 alone. Always layer PharmGKB level and a disproportionality readout, treating the three axes as orthogonal evidence streams.
- Treat FAERS-via-openFDA aggregate endpoints as a triage filter, not a verdict. Confirmatory DDI claims require report-level XML extracts.
- Account for indication confounding in pharmacovigilance signal interpretation (cyproheptadine / serotonin-syndrome co-reporting is the cautionary archetype here).
- For HLM platform validation, anchor on a hepatically-cleared reference set. IVIVE-direction signal on a mixed-clearance library will look weak even when the platform is performing correctly.
- Pharmacophore-level (not scaffold-level) analysis is the right granularity for SAR on chemically diverse hit sets.
Re-analysis of the GDPsm1 320-compound public panel from Ginkgo Datapoints. Methods: Echo-MS / QQQ-6500 acoustic-ejection MS for kinetic solubility, HLM Cl_int, and CYP2D6 IC50 with quinidine reference; Hill-fit IC50 quantitation on imax ≥ 50% curves; HLM literature comparison vs Obach 1999, Ito & Houston 2005, Shibata 2002 anchors; Fisher’s exact + BH FDR for substructure enrichment on Bemis–Murcko-deduplicated hits; openFDA FAERS via T6S2 with Bayesian Information Component (IC_lo95 > 0, Observed ≥ 3) as the disproportionality signal threshold; PharmGKB CPIC clinical-evidence levels (Hicks 2015, PMID 25974703; Hicks 2013, PMID 22205192); censoring flags retained for left-censored solubility, right-censored CYP2D6, and undetected stability values.
Interested in this kind of analysis?
See how Inflexa triangulates in-vitro ADME, clinical-evidence ontologies, and pharmacovigilance signals into translational-risk shortlists with full provenance.